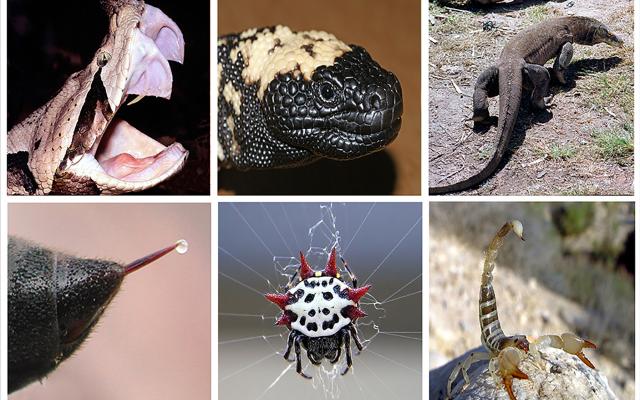
Animals as different as wasps and snakes have adopted surprisingly similar molecular mechanisms to squirt toxins out of their specialized cells. This is revealed by a study led by SIB scientists, who have conducted the first comparative analysis of gene expression profiles of venom glands. This result, published today in the Proceedings of the National Academy of Sciences, is particularly startling given that ‘venom factories’ have developed across different animals from scratch, rather than being inherited from a common ancestor.
Venomous animals represent about 15% of estimated global animal biodiversity. They are found in the deepest seas as well as on the driest land, and all have independently developed the ability to secrete venom. And it now seems that all venom factories use similar molecular mechanisms, whether what they produce merely creates an itching sensation, or causes total paralysis of the heart muscles.
A surprising molecular similarity across evolutionary distant animals
For essential organs with the same evolutionary origin (i.e. inherited from a common ancestor), such as human arms and bat wings, genetic similarity is very much expected. But venom glands, by contrast, are highly specialized structures that have evolved independently and sometimes recently across a range of animals. And whereas the human arm and the bat wing are both formed from the same tissue, venom glands arise from reproductive tissue in wasps as against mouthparts in snakes. Finally, venom factories are best known for their ability to secrete a cocktail of toxins which have some of the fastest evolving genes. For all these reasons, different molecular pathways could be expected to be involved in venom glands.
A surprising molecular similarity across evolutionary distant animals
For essential organs with the same evolutionary origin (i.e. inherited from a common ancestor), such as human arms and bat wings, genetic similarity is very much expected. But venom glands, by contrast, are highly specialized structures that have evolved independently and sometimes recently across a range of animals. And whereas the human arm and the bat wing are both formed from the same tissue, venom glands arise from reproductive tissue in wasps as against mouthparts in snakes. Finally, venom factories are best known for their ability to secrete a cocktail of toxins which have some of the fastest evolving genes. For all these reasons, different molecular pathways could be expected to be involved in venom glands.
Hence the surprise of the authors when they compared the genes expressed in 20 venomous species across the animal kingdom, from fishes to wasps and from scorpions to mammals. “What we found is a strong convergence in the global gene expression levels of venom glands across very different animals”, says SIB’s Giulia Zancolli, Marie Skłodowska-Curie postdoctoral fellow in the team led by Marc Robinson-Rechavi and Frédéric Bastian (UNIL). “It seems Nature has simply found the optimal way for cells to achieve their specific venom secreting-function.”
A large-scale gene expression comparison across 20 species with SIB software tools and databases
Among other steps, the scientists had to group the protein sequences from the species selected into venom-related ‘orthogroups’, i.e. sets of orthologous genes across all species. They did this grouping using OrthoDB, and protein function hints from UniProtKB/SwissProt. To find the anatomical entities in which tissue-specific gene modules were enriched, they compared the gene expression levels reflected by publicly-available RNAseq datasets, and relied on the TopAnat tool of the Bgee resource for gene expression pattern.
What are the next research steps envisaged?
Despite the widespread presence of venom systems across the animal kingdom, and the medical focus on the toxins they produce, we still know very little about their origins and genetic basis. The next research steps will be to fill in gaps in knowledge of the genetic mechanisms of the emergence and development of toxins. “This study represents the first step towards an understanding of the molecular mechanisms underlying the repeated evolution of one of the most successful adaptive traits in the animal kingdom”, concludes Zancolli.
Reference(s)
Zancolli G et al. Convergent evolution of venom gland transcriptomes across Metazoa. PNAS, 2022.