Today, the delta variant of the virus dominates the Swiss landscape. This is only possible thanks to an unprecedented sequencing effort. At the heart of the process, a national infrastructure, co-led by the SIB Swiss Institute of Bioinformatics, now centralises all the genetic sequences of the virus collected in Switzerland. On the one hand, it provides the Federal Office of Public Health (FOPH) with an automated overview of the distribution and emergence of variants throughout the country. On the other hand, it allows the rapid and mass sharing of Swiss sequences with international databases. This places Switzerland among the largest contributors of SARS-CoV-2 sequences worldwide, thereby accelerating research into vaccines and treatments.
About the Swiss Pathogen Surveillance Platform, SPSP
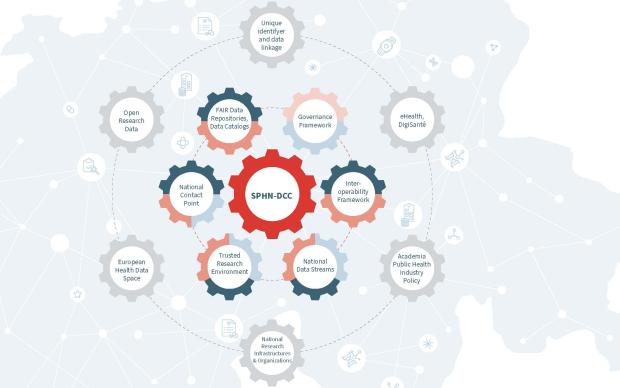
SPSP.ch is a collaborative platform following the Onehealth approach, i.e. multidisciplinary and aiming, among other things, to optimise human health outcomes. It is co-managed by the SIB in collaboration with the University Hospitals of Basel, Lausanne and Geneva, as well as the Universities of Bern and Zurich (see the complete list of partners contributing data). It is hosted on the secure IT infrastructure of the SIB and complies with the data security standards of the Swiss Personalised Health Network (SPHN).
The need for coordinated monitoring of variants at the national level
Alpha yesterday, Delta today and Mu perhaps tomorrow: Switzerland is on the lookout for the appearance of new variants and their transmission chains. The sequencing of the virus genome makes it possible to determine, from a positive PCR test, the identity of the variant concerned and its complete genetic profile. In August 2021 alone, more than 5,600 sequences were analysed by more than ten academic or private laboratories throughout Switzerland. But these data only make sense if they are related to each other as quickly as possible - hence the essential need for coordination and standardisation.
This is where a collaborative and secure infrastructure co-led by the SIB comes in. Its name? The Swiss Pathogen Surveillance Platform (SPSP). Its role? To centralise, harmonise and annotate SARS-CoV-2 genomic sequences from laboratories in Switzerland, as well as the accompanying data (date of PCR test, sampling method, reason for sequencing, machine used, location of test, gender and age of patient). Its aim? To support the national genomic surveillance strategy conducted jointly by the National Reference Centre for Emerging Viral Infections (CRIVE) and the FOPH. The platform allows the authorities to benefit from a global and automated view of sequencing in Switzerland, and feeds international databases used for research on the virus.
"So far, the sequences we have received come from almost all Swiss cantons: this is excellent news, as it means that new variants are unlikely to fly under the radar," explains Aitana Lebrand, Team Lead Data Science at SIB in charge of the SPSP platform.
Accelerating the monitoring of the epidemic in Switzerland
Three times a week, the platform sends its genomic surveillance report to the FOPH. The FOPH integrates it into its statistics, and can also cross-check the information with the patient data it has on hospitalisations, vaccination, symptoms at the time of the test, etc. This could make it possible to identify that one mutation among all those observed seems to be linked to a higher pathogenicity, or higher resistance to vaccines.
"Thanks to the surveillance platform developed by SIB, we can access a centralised and standardised database through a single entry point, rather than receiving reports from each laboratory in different formats," explains Mirjam Mäusezahl, co-director of the epidemiology section at the FOPH. « For us, this represents a huge time saving, as well as an increased granularity in the analysis of the sequencing data. It allows us to focus our efforts on adapting public health policies. »
Boosting international research by facilitating open science
With funding from the State Secretariat for Education, Research and Innovation SERI, the SPSP platform also transmits fully anonymised viral sequences to open science platforms, such as the European COVID-19 portal, to boost international research. Thanks to these efforts, Switzerland currently ranks 4th in terms of the number of shared SARS-CoV-2 sequences, after the UK, the US and Germany. These public databases are fundamental for studying and understanding the role of the observed variations on the pathogenicity of the virus, its interactions with host cells at the time of infection, and for the development of vaccines and treatments
What's next? A national SPSP application, beyond COVID-19
This collaboration with the FOPH in the context of the current pandemic is in line with the initial mission of the SIB platform: to enable specialists to rapidly identify the emergence and spread of pathogens, and to take early measures to contain transmission by tracking them in near real time. Before COVID-19 changed research priorities, SPSP specifically targeted multi-resistant bacteria. "In the age of sequencing, this tool meets an essential need for microbiologists to have a harmonised and centralised analysis of molecular profiles of infectious agents on a national scale," comments Adrian Egli, Head of the Department of Bacteriology and Mycology at the University Hospital of Basel and co-leader of the platform.
"The possibility of using SPSP in the long term to link genomic data of emerging bacteria or viruses in Switzerland with epidemiological data is promising to ensure exemplary responsiveness of the country in the field of public health," concludes Mirjam Mäusezahl.
Read the press release in French and German
Selected press coverage: Le Temps, Heidi.news, RTS La 1ère CQFD, Tages Anzeiger